




















中国农业科技导报 ›› 2024, Vol. 26 ›› Issue (6): 11-21.DOI: 10.13304/j.nykjdb.2024.0172
鲍新跃1,2( ), 陈红敏3(
), 陈红敏3( ), 王伟伟2, 唐益苗2, 房兆峰2, 马锦绣2, 汪德州2(
), 王伟伟2, 唐益苗2, 房兆峰2, 马锦绣2, 汪德州2( ), 左静红2(
), 左静红2( ), 姚占军1(
), 姚占军1( )
)
收稿日期:2024-03-07
接受日期:2024-03-26
出版日期:2024-06-15
发布日期:2024-06-12
通讯作者:
汪德州,左静红,姚占军
作者简介:鲍新跃E-mail:1772145342@qq.com基金资助:
Xinyue BAO1,2( ), Hongmin CHEN3(
), Hongmin CHEN3( ), Weiwei WANG2, Yimiao TANG2, Zhaofeng FANG2, Jinxiu MA2, Dezhou WANG2(
), Weiwei WANG2, Yimiao TANG2, Zhaofeng FANG2, Jinxiu MA2, Dezhou WANG2( ), Jinghong ZUO2(
), Jinghong ZUO2( ), Zhanjun YAO1(
), Zhanjun YAO1( )
)
Received:2024-03-07
Accepted:2024-03-26
Online:2024-06-15
Published:2024-06-12
Contact:
Dezhou WANG,Jinghong ZUO,Zhanjun YAO
摘要:
小麦产量关系我国粮食安全,干旱、低温、盐害和高温等非生物胁迫严重制约小麦产量增长。前期转录组分析发现小麦TaCOBL-5D在多种非生物胁迫下差异表达。克隆并获取TaCOBL-5D及其同源基因TaCOBL-5A、TaCOBL-5B,并对其生物信息学特性及表达模式进行了分析。结果显示,TaCOBL-5与其他物种的COBL基因在基因结构、蛋白三级结构、保守结构域以及启动子调控元件方面表现出明显的保守性。TaCOBL-5在根部表达量最高,并对各种非生物胁迫响应不同,尤其是在干旱胁迫下调控显著,表明其在干旱胁迫中的重要性,同时对低温、高温和盐胁迫也有不同响应。此外,TaCOBL-5D基因在不同干旱抗性及高温抗性材料中表达量差异显著,进一步暗示其在逆境胁迫中具有重要作用。这些研究结果有助于理解COBL基因在小麦中的功能,同时为小麦抗逆育种提供科学支持。
中图分类号:
鲍新跃, 陈红敏, 王伟伟, 唐益苗, 房兆峰, 马锦绣, 汪德州, 左静红, 姚占军. 小麦TaCOBL-5基因克隆及表达分析[J]. 中国农业科技导报, 2024, 26(6): 11-21.
Xinyue BAO, Hongmin CHEN, Weiwei WANG, Yimiao TANG, Zhaofeng FANG, Jinxiu MA, Dezhou WANG, Jinghong ZUO, Zhanjun YAO. Cloning and Expression Analysis of Wheat TaCOBL-5 Genes[J]. Journal of Agricultural Science and Technology, 2024, 26(6): 11-21.
引物名称 Primer name | 正向引物序列 Forward primer sequence (3’-5’) | 反向引物序列 Reverse primer sequence (3’-5’) |
|---|---|---|
| TaCOBL-5A | GCGGCACCCGTGTCTTCTAT | CGTCTCGTCTCGTCGCAGTA |
| TaCOBL-5B | GCGGCACCCATGTCTTCTAT | CGTCTCGTCTCGTCGCAGTA |
| TaCOBL-5D | ACGGCACCCGCGTCTTCTAT | TCTCGTCGCTGTAAAAACTG |
| qTaCOBL-5A | CGTTGGATCTCTCTTGCAGC | TGGGATGGTCATGGGCAAAG |
| qTaCOBL-5B | GATTACGTGCAGGTTACATTCC | TCTCAAGGCTCCAGGTCAGG |
| qTaCOBL-5D | CAGCGAATCATAAGCCTCTG | GAGTAGCGGGGCAGGAAATG |
| TaActin | GGAATCCATGAGACCACCTAC | GACCCAGACAACTCGCAAC |
表1 基因克隆和荧光定量的引物序列
Table 1 Primers used in this study for gene cloning and qPCR
引物名称 Primer name | 正向引物序列 Forward primer sequence (3’-5’) | 反向引物序列 Reverse primer sequence (3’-5’) |
|---|---|---|
| TaCOBL-5A | GCGGCACCCGTGTCTTCTAT | CGTCTCGTCTCGTCGCAGTA |
| TaCOBL-5B | GCGGCACCCATGTCTTCTAT | CGTCTCGTCTCGTCGCAGTA |
| TaCOBL-5D | ACGGCACCCGCGTCTTCTAT | TCTCGTCGCTGTAAAAACTG |
| qTaCOBL-5A | CGTTGGATCTCTCTTGCAGC | TGGGATGGTCATGGGCAAAG |
| qTaCOBL-5B | GATTACGTGCAGGTTACATTCC | TCTCAAGGCTCCAGGTCAGG |
| qTaCOBL-5D | CAGCGAATCATAAGCCTCTG | GAGTAGCGGGGCAGGAAATG |
| TaActin | GGAATCCATGAGACCACCTAC | GACCCAGACAACTCGCAAC |
基因 Gene | 基因号 Gene ID number | 物理位置 Physical position/bp | 分子量 Molecular weight/Da | 等电点 pI | 蛋白长度 Protein length/aa | 预测定位 Predicted location |
|---|---|---|---|---|---|---|
| TaCOBL-5A | TraesCS5A02G392000 | 588 375 577~588 379 139 | 50 867.2 | 8.98 | 457 | 细胞膜Cell membrane |
| TaCOBL-5B | TraesCS5B02G396900 | 574 675 118~574 678 654 | 50 841.18 | 8.98 | 457 | 细胞膜Cell membrane |
| TaCOBL-5D | TraesCS5D02G401900 | 467 600 940~467 604 428 | 50 823.15 | 8.98 | 457 | 细胞膜Cell membrane |
表2 TaCOBL-5基因信息及其编码蛋白的理化性质分析
Table. 2 TaCOBL-5 gene information and physicochemical properties analysis
基因 Gene | 基因号 Gene ID number | 物理位置 Physical position/bp | 分子量 Molecular weight/Da | 等电点 pI | 蛋白长度 Protein length/aa | 预测定位 Predicted location |
|---|---|---|---|---|---|---|
| TaCOBL-5A | TraesCS5A02G392000 | 588 375 577~588 379 139 | 50 867.2 | 8.98 | 457 | 细胞膜Cell membrane |
| TaCOBL-5B | TraesCS5B02G396900 | 574 675 118~574 678 654 | 50 841.18 | 8.98 | 457 | 细胞膜Cell membrane |
| TaCOBL-5D | TraesCS5D02G401900 | 467 600 940~467 604 428 | 50 823.15 | 8.98 | 457 | 细胞膜Cell membrane |
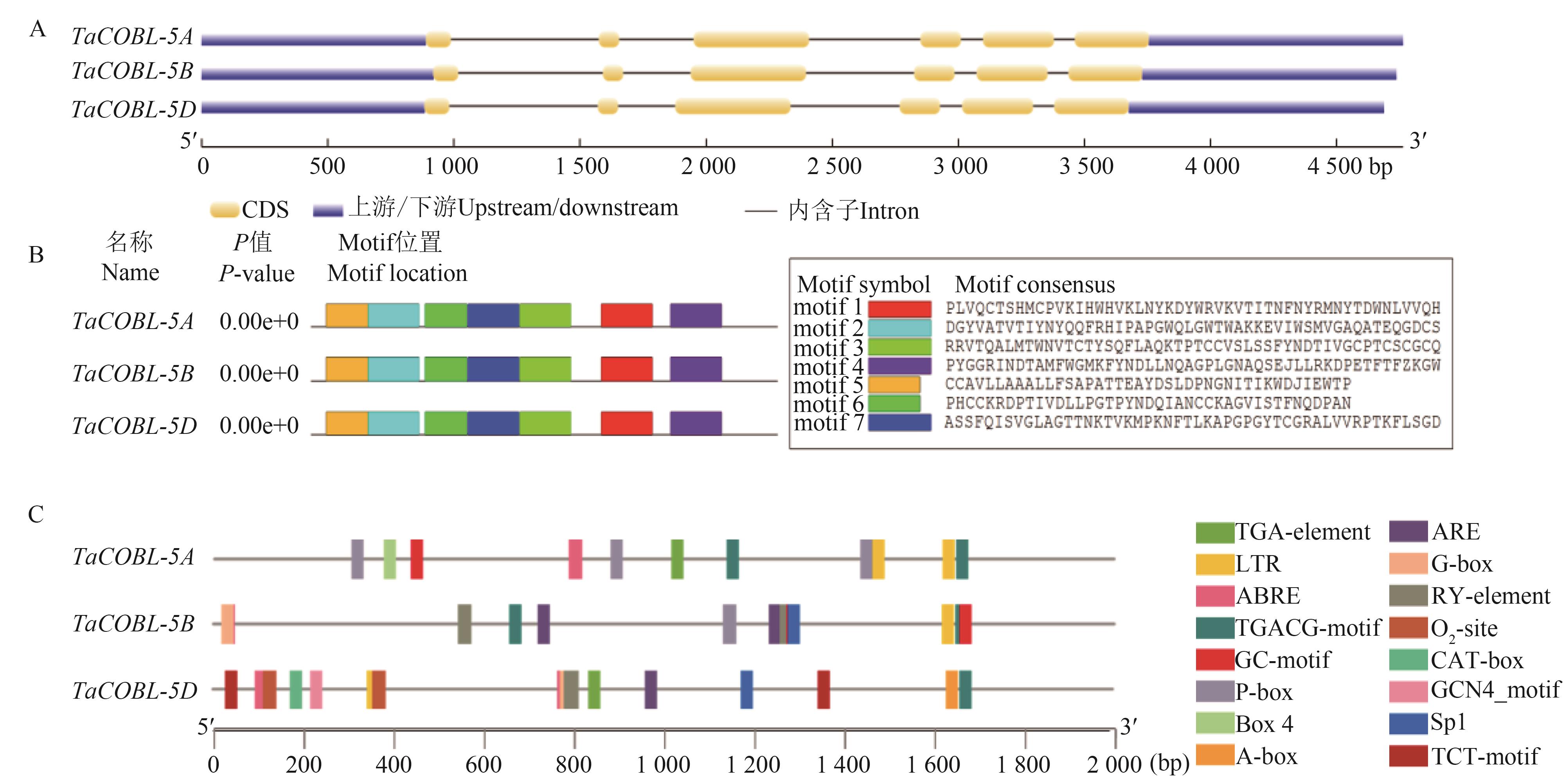
图1 TaCOBL-5基因结构、保守基序及启动子顺式作用元件分析A:基因结构;B:保守序列;C:启动子顺式作用元件
Fig. 1 Analysis of gene structure, conserved motif and cis-acting regulatory elements of TaCOBL-5A: Gene structure; B: Conserved motif; C: Cis-acting regulatory elements
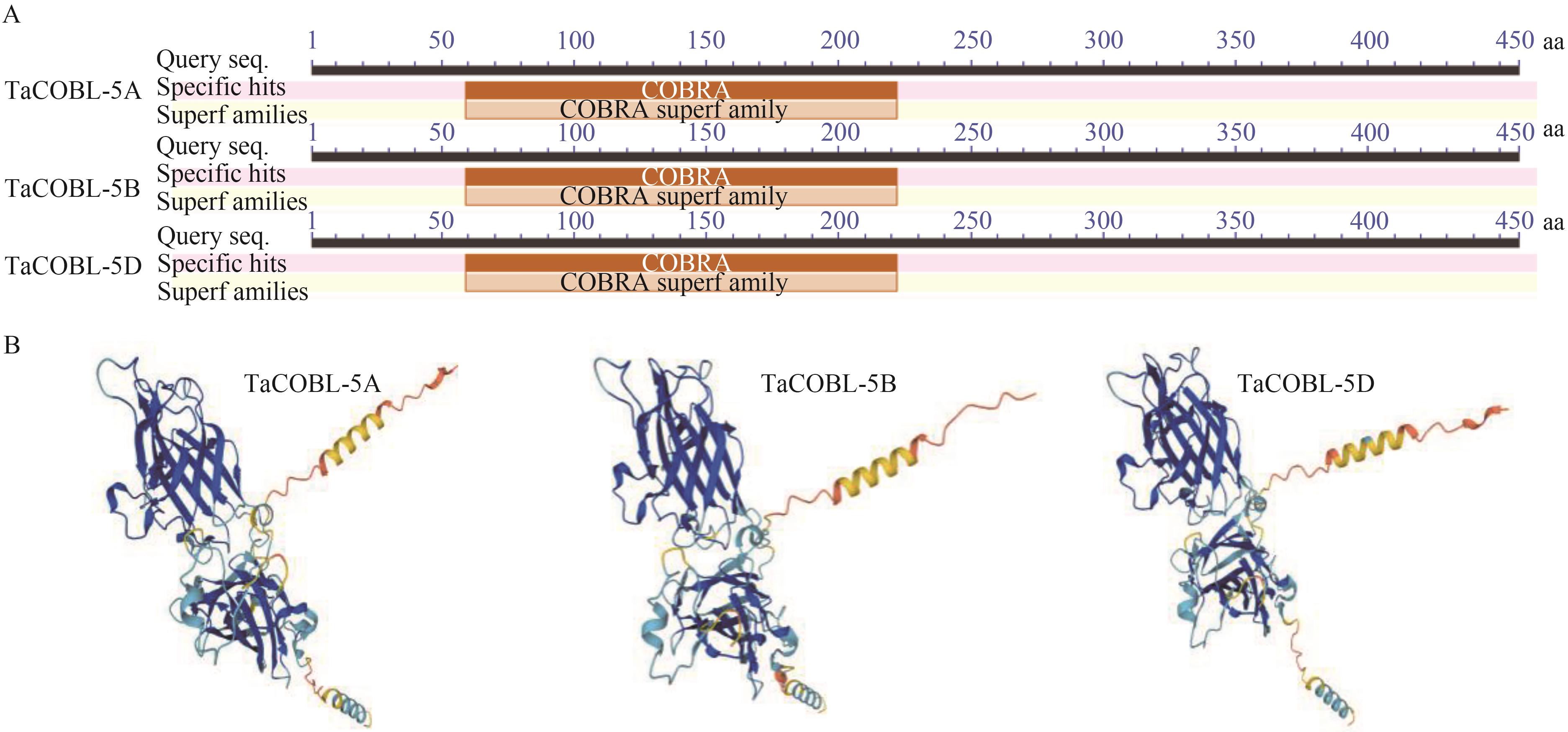
图3 COBRA结构域及三级结构预测A: COBRA结构域;B:COBL蛋白三级结构
Fig. 3 Analysis of conserved COBRA domain and the tertiary structure of COBL proteinsA: COBRA domain; B: Tertiary structure of COBL protein
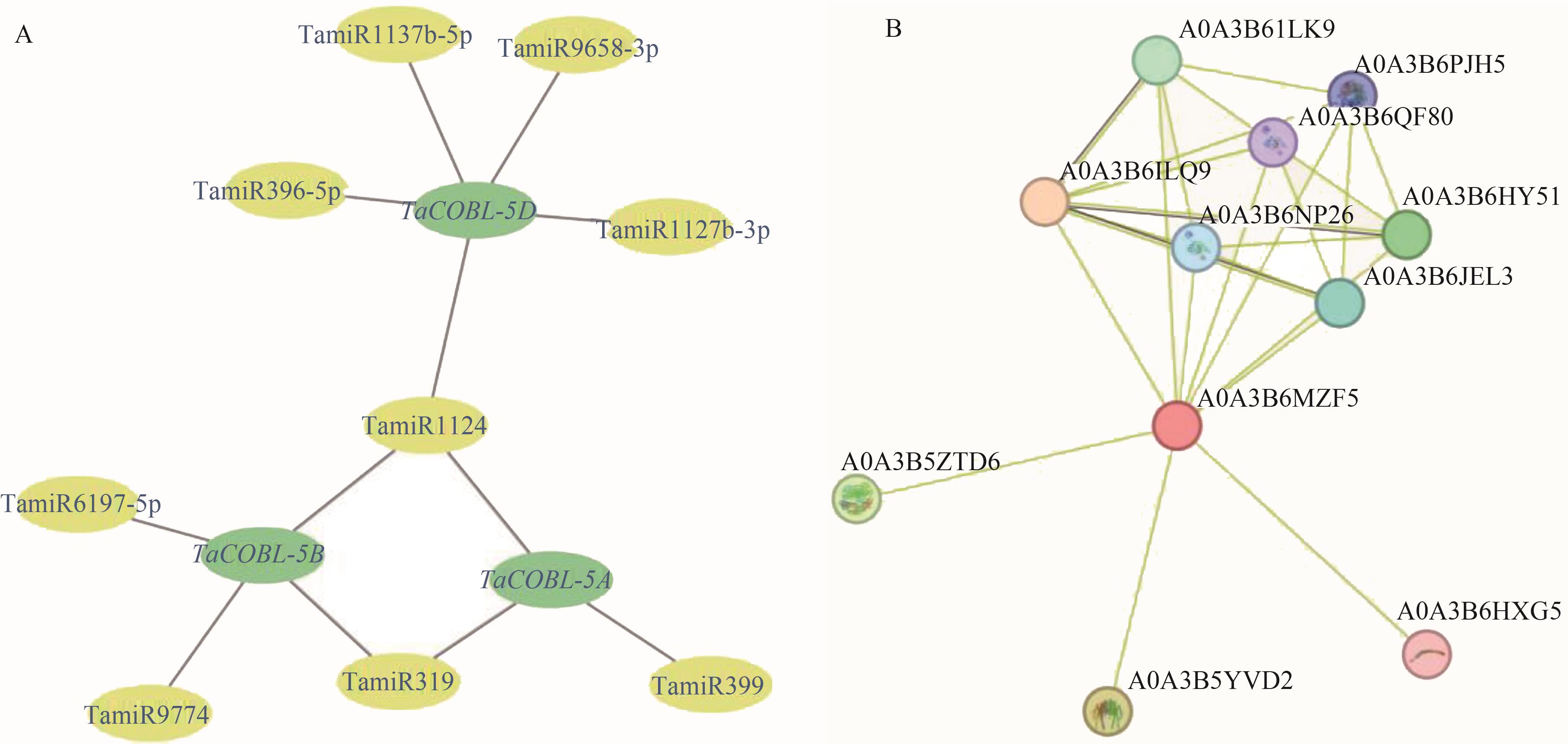
图5 小麦miRNA调控网络预测及互作蛋白预测A: miRNA调控网络;B:蛋白互作网络
Fig. 5 Prediction putative networks of wheat miRNAs and the interacting proteinsA: Putative network of wheat miRNAs; B: Network of interacting proteins
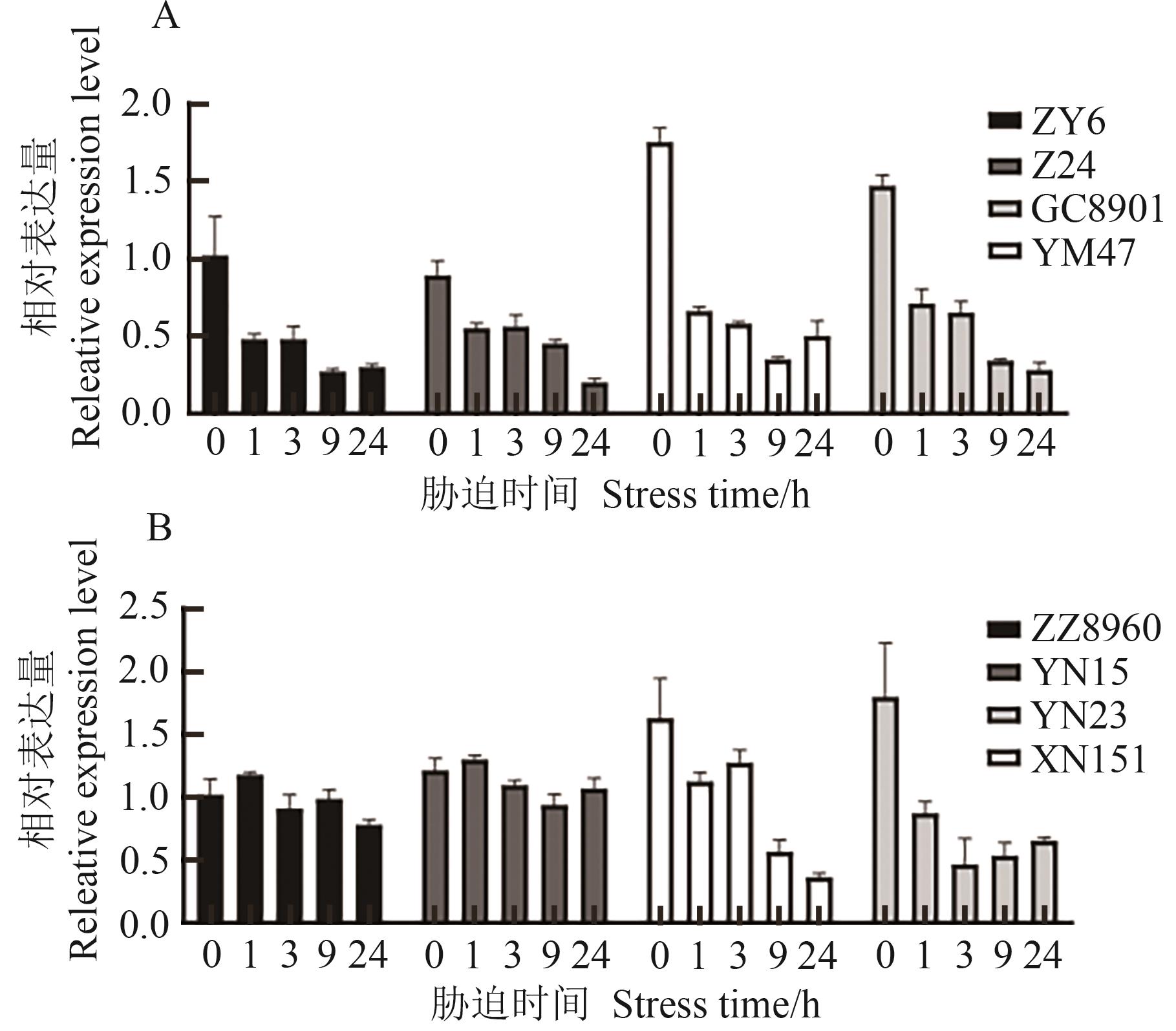
图8 不同抗性材料干旱胁迫、高温胁迫处理下TaCOBL-5D表达模式分析A: 干旱胁迫;B:高温胁迫
Fig. 8 Expression profiles of TaCOBL-5D among different resistant materials under drought and heat stressesA: Drought stress; B: Heat stress
| 1 | ZHU J K. Salt and drought stress signal transduction in plants [J]. Annu. Rev. Plant Biol., 2002, 53(1):247-273. |
| 2 | 陈翔,胡雨喆,陈甜甜,等.小麦抗低温逆境化控技术研究进展[J].植物营养与肥料学报, 2023, 29(8):1543-1555. |
| CHEN X, HU Y Z, CHEN T T, et al.. Progress of chemical regulation on wheat resistance to low temperature stress [J]. J. Plant Nutr. Fert., 2023, 29(8):1543-1555. | |
| 3 | WINFIELD M O, LU C G, WILSON I D, et al.. Plant responses to cold: transcriptome analysis of wheat [J]. Plant Biotechnol. J., 2010, 8(7):749-771. |
| 4 | MAO H D, JIAN C, CHENG X X, et al.. The wheat ABA receptor gene TaPYL1-1B contributes to drought tolerance and grain yield by increasing water-use efficiency [J]. Plant Biotechnol. J., 2022, 20(5):846-861. |
| 5 | 温辉芹,程天灵,裴自友,等.山西中部区试小麦品种抗旱节水指标分析[J].山西农业科学,2020,48(10):1572-1575. |
| WEN H Q, CHENG T L, PEI Z Y, et al.. Analysis on drought resistance and water saving indexes of wheat varieties of regional trial in central Shanxi [J]. J. Shanxi Agric. Sci., 2020, 48(10):1572-1575. | |
| 6 | 健康,倪建平.植物非生物胁迫信号转导及应答[J].中国稻米, 2016, 22(6):52-60. |
| ZHU J K, NI J P. Abiotic stress signaling and responses in plants [J]. China Rice, 2016, 22(6):52-60. | |
| 7 | 盛松柏,田菊,庞晓明.冬枣COBRA基因家族全基因组鉴定及表达分析[J].分子植物育种, 2018, 16(1):61-68. |
| SHENG S B, TIAN J, PANG X M. Genome-wide identification and expression analysis of COBRA gene family in Ziziphus jujuba [J]. Mol. Plant Breeding, 2018, 16(1):61-68. | |
| 8 | SUN X M, XIONG H Y, JIANG C H, et al.. Natural variation of DROT1 confers drought adaptation in upland rice [J/OL]. Nat. Commun., 2022, 13(1):4265 [2024-04-03].. |
| 9 | LI Y H, QIAN O, ZHOU Y H, et al.. BRITTLE CULM1, which encodes a COBRA-like protein, affects the mechanical properties of rice plants [J]. Plant Cell, 2003, 15(9):2020-2031. |
| 10 | BRADY S M, SONG S, DHUGGA K S, et al.. Combining expression and comparative evolutionary analysis. The COBRA gene family [J]. Plant Physiol., 2007, 143(1):172-187. |
| 11 | JULIUS B T, MCCUBBIN T J, MERTZ R A, et al.. Maize Brittle Stalk2-Like3, encoding a COBRA protein, functions in cell wall formation and carbohydrate partitioning [J]. Plant Cell, 2021, 33(10):3348-3366. |
| 12 | AHMED M Z, ALQAHTANI A S, NASR F A, et al.. Comprehensive analysis of the COBRA-like (COBL) gene family through whole-genome analysis of land plants [J]. Genet. Resour. Crop Evol., 2024, 71(1):863-872. |
| 13 | QIU C, CHEN J H, WU W H, et al.. Genome-wide analysis and abiotic stress-responsive patterns of COBRA-like gene family in Liriodendron chinense [J/OL]. Plants-Basel, 2023, 12(8):1616 [2024-04-03]. . |
| 14 | BEN-TOV D, ABRAHAM Y, STAV S, et al.. COBRA-LIKE2, a member of the glycosylphosphatidylinositol-anchored COBRA-LIKE family, plays a role in cellulose deposition in Arabidopsis seed coat mucilage secretory cells [J]. Plant Physiol., 2015, 167(3):711-724. |
| 15 | LIU F F, WAN Y X, CAO W X, et al.. Novel function of a putative TaCOBL ortholog associated with cold response [J]. Mol. Biol. Rep., 2023, 50(5):4375-4384. |
| 16 | LIU L F, SHANG-GUAN K K, ZHANG B C, et al.. Brittle Culm1, a COBRA-Like protein, functions in cellulose assembly through binding cellulose microfibrils [J/OL]. PLoS Genet., 2013, 9(8):e1003704 [2024-04-03]. . |
| 17 | YE X, KANG B G, OSBURN D L, et al.. The COBRA gene family in Populus and gene expression in vegetative organs and in response to hormones and environmental stresses [J]. Plant Growth Regul., 2009, 58(2):211-223. |
| 18 | ZHANG D Q, YANG X H, ZHANG Z Y, et al.. Expression and nucleotide diversity of the poplar COBL gene [J]. Tree Genet. Genomes, 2010, 6(2):331-344. |
| 19 | YILAN E, XIN G, JING X, et al.. Genome-wide identification of the COBRA-like gene family in Pinus tabuliformis and the role of PtCOBL12 in the regulation of cellulose biosynthesis [J]. Ind. Crop Prod., 2023, 203. |
| 20 | YANG Q, WANG S, CHEN H, et al.. Genome-wide identification and expression profiling of the COBRA-like genes reveal likely roles in stem strength in rapeseed (Brassica napus L.) [J/OL]. PLoS One, 2021,16(11):e0260268 [2024-04-03]. . |
| 21 | NIU E L, SHANG X G, CHENG C Z, et al.. Comprehensive analysis of the COBRA-Like (COBL) gene family in Gossypium identifies two COBLs potentially associated with fiber quality [J/OL]. PLoS One, 2015, 10(12):e014572 [2024-04-03]. . |
| 22 | SANGI S, ARAÚJO PAULA M, COELHO F S, et al.. Genome-wide analysis of the COBRA-Like gene family supports gene expansion through whole-genome duplication in soybean (Glycine max) [J]. Plants, 2021, 10 (1):167-167. |
| 23 | XU L, WANG D Z, LIU S, et al.. Comprehensive atlas of wheat (Triticum aestivum L.) AUXIN RESPONSE FACTOR expression during male reproductive development and abiotic stress [J/OL]. Front. Plant Sci., 2020, 11:586144 [2024-04-03]. . |
| 24 | 吴凯铭.337份小麦品种(系)萌发期抗旱性鉴定及生理响应[D].杨凌:西北农林科技大学,2022. |
| WU K M. Identification of drought resistance and physiological response during germination in 337 wheat varieties (lines) [D]. Yangling: Northwest A&F University, 2022. | |
| 25 | 史冰新.黄淮和长江中下游冬麦区小麦耐热种质资源筛选[D].杨凌:西北农林科技大学,2023. |
| SHI B X. Screened for heat tolerance of wheat germplasm resources in the Yellow and Huai River valley winter wheat zone and the middle and lower Yangtze valley winter wheat zone [D]. Yangling: Northwest A&F University, 2023. | |
| 26 | KESAWAT M S, KHERAWAT B S, SINGH A, et al.. Genome-wide identification and characterization of the Brassinazole-resistant (BZR) gene family and its expression in the various developmental stage and stress conditions in wheat (Triticum aestivum L.) [J]. Int. J. Mol. Sci., 2021, 22(16):8743-8743. |
| 27 | FANG Y J, ZHENG Y Q, LU W, et al.. Roles of miR319-regulated TCPs in plant development and response to abiotic stress [J]. Crop J., 2021, 9(1):17-28. |
| 28 | JIAN C, HAO P A, HAO C Y, et al.. The miR319/TaGAMYB3 module regulates plant architecture and improves grain yield in common wheat (Triticum aestivum) [J]. New Phytol., 2022, 235(4):1515-1530. |
| 29 | 梁婷,左静红,陆青,等.小麦IQM基因家族鉴定及非生物胁迫下表达分析[J].中国农业科技导报, 2023, 25(2):27-37. |
| LIANG T, LU Q, ZUO J Het al.. Identification and expression analysis under abiotic stress of IQM gene family in wheat (Triticum aestivum L.) [J]. J. Agric. Sci. Technol., 2023, 25(2):27-37. | |
| 30 | 陆青,梁婷,汪德州,等.小麦热激蛋白基因TaHSP90-1的克隆与表达分析[J].中国农业科技导报, 2022, 24(8):44-54. |
| LU Q, LIANG T, WANG D Zet al.. Cloning and expression analysis of wheat heat shock protein gene TaHSP90-1 [J]. J. Agric. Sci. Technol., 2022, 24(8):44-54. |
| [1] | 刘一凡, 刘少帅, 臧瑞, 李洋, 刘薇, 李婷婷, 刘旦梅, 刘登才, 李爱丽, 毛龙, 王翔, 耿帅锋. 168份小麦种质资源品质性状分析[J]. 中国农业科技导报, 2025, 27(9): 44-57. |
| [2] | 吕彩霞, 李永福, 信会男, 李娜, 赖宁, 耿庆龙, 陈署晃. 缓释氮肥对滴灌冬小麦产量及土壤硝/铵态氮的影响[J]. 中国农业科技导报, 2025, 27(8): 179-186. |
| [3] | 杨志宁, 陆许可, 范亚朋, 孙玉平, 喻欣, 王亮, 高永健, 古丽加依那什·哈不力, 古丽沙西·拿斯依, 吾木尔·阿不都, 黄慧, 张梦豪, 王立东, 陈筱, 肖磊, 张鑫蕊, 王帅, 陈修贵, 王俊娟, 郭丽雪, 高文伟, 叶武威. 棉花GhDMT7基因对5-氮杂胞嘧啶核苷的响应机制[J]. 中国农业科技导报, 2025, 27(8): 28-35. |
| [4] | 朱强, 车宗贤, 崔恒, 张久东, 包兴国. 绿肥替代氮肥对麦田温室气体的影响[J]. 中国农业科技导报, 2025, 27(7): 182-189. |
| [5] | 胡懿, 公杰, 赵玮, 程蓉, 柳忠玉, 高世庆, 杨亚珍. 小麦PHY基因家族鉴定及热胁迫下表达分析[J]. 中国农业科技导报, 2025, 27(7): 30-43. |
| [6] | 孙吉林, 张家琦, 孟凡森, 孙思龙. 二倍体长穗偃麦草HSP70基因家族鉴定及表达模式分析[J]. 中国农业科技导报, 2025, 27(6): 28-38. |
| [7] | 呼斯乐, 包玉龙, 图布新巴雅尔null, 陶际峰, 郭恩亮. 基于无人机高光谱和集成学习的春小麦叶绿素含量反演[J]. 中国农业科技导报, 2025, 27(6): 93-103. |
| [8] | 史硕, 冯宇, 李亮, 孟瑞, 章延泽, 杨秀荣. 印度梨形孢介导小麦抗纹枯病的转录组分析及关键基因筛选[J]. 中国农业科技导报, 2025, 27(5): 133-145. |
| [9] | 马蓓, 公杰, 杜银柯, 甘雨薇, 程蓉, 朱波, 易丽霞, 马锦绣, 高世庆. 小麦花粉孔发育相关TaINP1基因鉴定及表达分析[J]. 中国农业科技导报, 2025, 27(4): 22-35. |
| [10] | 陈宜新, 杨秀波, 田士军, 王聪, 白志英, 李存东, 张科. 陆地棉GhCOMT28对干旱胁迫的响应[J]. 中国农业科技导报, 2025, 27(4): 45-56. |
| [11] | 刘金仙, 王丽娟, 刘杰, 傅仙玉, 武广珩. 茶树钙调蛋白结合转录激活因子(CAMTA)家族基因鉴定与表达分析[J]. 中国农业科技导报, 2025, 27(3): 71-82. |
| [12] | 薛振宇, 张康康, 张元元, 闫强强, 姚立蓉, 张宏, 孟亚雄, 司二静, 李葆春, 马小乐, 王化俊, 汪军成. 优质抗旱小麦种质的筛选及功能基因检测[J]. 中国农业科技导报, 2025, 27(1): 35-49. |
| [13] | 郭肖蓉, 刘颖, 樊佳珍, 黄涛, 周荣. 猪CREBRF基因生物信息学和表达规律分析[J]. 中国农业科技导报, 2024, 26(9): 44-53. |
| [14] | 孙宪印, 牟秋焕, 米勇, 吕广德, 亓晓蕾, 孙盈盈, 尹逊栋, 王瑞霞, 吴科, 钱兆国, 赵岩, 高明刚. 基于GT双标图对小麦新品系的分类评价[J]. 中国农业科技导报, 2024, 26(7): 14-24. |
| [15] | 陈明迪, 胡桂花, 张海文, 王旺田. 水稻RR基因家族生物信息学及表达模式分析[J]. 中国农业科技导报, 2024, 26(5): 20-29. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||